
Khoi Bui Dinh
Past and on-going projects

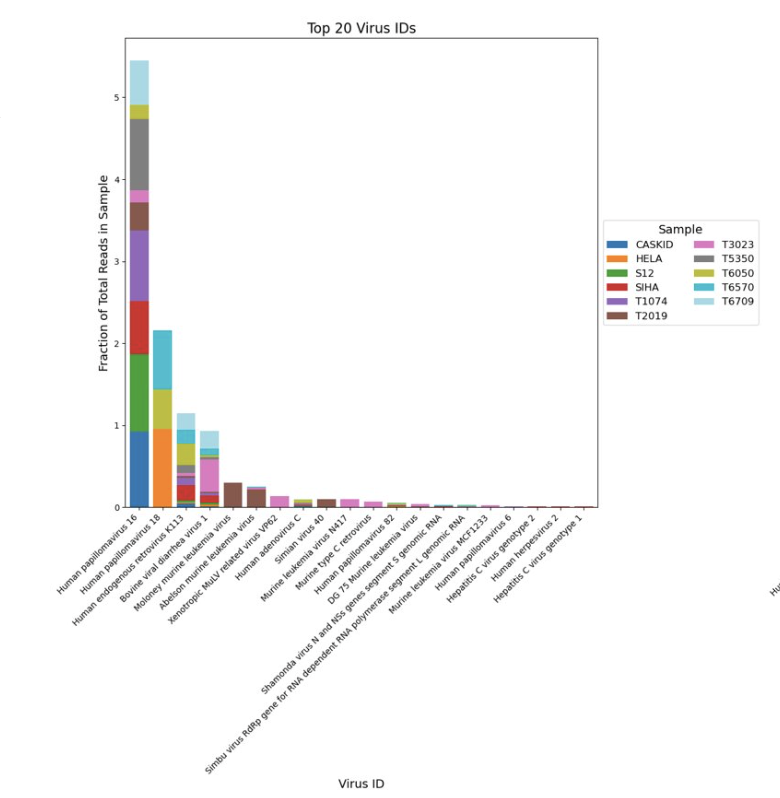
Viral detection in public human RNAseq data
Tools:
nextflow
nf-core
R
Abstract:
Cervical cancer is one of the leading causes of death in women, ranking as the fourth most common cancer type worldwide. Human papillomavirus (HPV) is known to be a driver in causing cervical cells to grow abnormally. Yet, in human transcriptomics sequencing, most of the reads unaligned to the human reference are usually discarded even though they may contain useful taxonomic and viral expression information. Here we propose that by developing a workflow to preserve and analyze viral reads found in human samples, viral strains identification can support diagnostic potential in virus-associated human cancers. We aligned dehosted reads from human cervical cancer to a curated viral database, showing that there are differences in gene expression between HPV-specific infected samples using public datasets. We used the gene expression matrix as features for model training, and found that viral reads can potentially be used for classification tasks.
Cervical cancer is one of the leading causes of death in women, ranking as the fourth most common cancer type worldwide. Human papillomavirus (HPV) is known to be a driver in causing cervical cells to grow abnormally. Yet, in human transcriptomics sequencing, most of the reads unaligned to the human reference are usually discarded even though they may contain useful taxonomic and viral expression information. Here we propose that by developing a workflow to preserve and analyze viral reads found in human samples, viral strains identification can support diagnostic potential in virus-associated human cancers. We aligned dehosted reads from human cervical cancer to a curated viral database, showing that there are differences in gene expression between HPV-specific infected samples using public datasets. We used the gene expression matrix as features for model training, and found that viral reads can potentially be used for classification tasks.

Responsible AI for Genomics Research - A Review
Tools:
shap
lime
pytorch
Abstract:
Artificial intelligence (AI) has become a crucial tool in genomics research, driven by advances in high-throughput sequencing technologies that generate vast amounts of data. While AI enables the integration and interpretation of diverse omics data, it also raises concerns about trust and responsibility. As AI systems grow more complex, the demand for Responsible and Explainable AI (XAI), which enhances the transparency of AI outputs and decision-making, has emerged as a critical priority in ensuring ethical use. We conducted a comprehensive literature review to select 23 AI models applicable to various fields within genomics, including synthetic biology and cancer research. These models were evaluated based on the FAIR principles: Findable, Accessible, Interoperable, and Reusable. Our findings reveal that while the models scored between 60% and 90% across all metrics, they particularly excelled in Findable and Accessible. However, their Interoperable scores were lower, indicating a lack of design for broad application in diverse workflows, which limits their potential impact. This study highlights the need for responsible and FAIR practices in AI model development for genomics. We recommend focusing on Interoperability to enhance the usability of models across different settings, ensuring reproducibility through containerization, and facilitating future enhancements by maintaining accessible code repositories. By adopting these practices, the AI model authors will be able to replicate and extend the impacts of their AI models.
Artificial intelligence (AI) has become a crucial tool in genomics research, driven by advances in high-throughput sequencing technologies that generate vast amounts of data. While AI enables the integration and interpretation of diverse omics data, it also raises concerns about trust and responsibility. As AI systems grow more complex, the demand for Responsible and Explainable AI (XAI), which enhances the transparency of AI outputs and decision-making, has emerged as a critical priority in ensuring ethical use. We conducted a comprehensive literature review to select 23 AI models applicable to various fields within genomics, including synthetic biology and cancer research. These models were evaluated based on the FAIR principles: Findable, Accessible, Interoperable, and Reusable. Our findings reveal that while the models scored between 60% and 90% across all metrics, they particularly excelled in Findable and Accessible. However, their Interoperable scores were lower, indicating a lack of design for broad application in diverse workflows, which limits their potential impact. This study highlights the need for responsible and FAIR practices in AI model development for genomics. We recommend focusing on Interoperability to enhance the usability of models across different settings, ensuring reproducibility through containerization, and facilitating future enhancements by maintaining accessible code repositories. By adopting these practices, the AI model authors will be able to replicate and extend the impacts of their AI models.
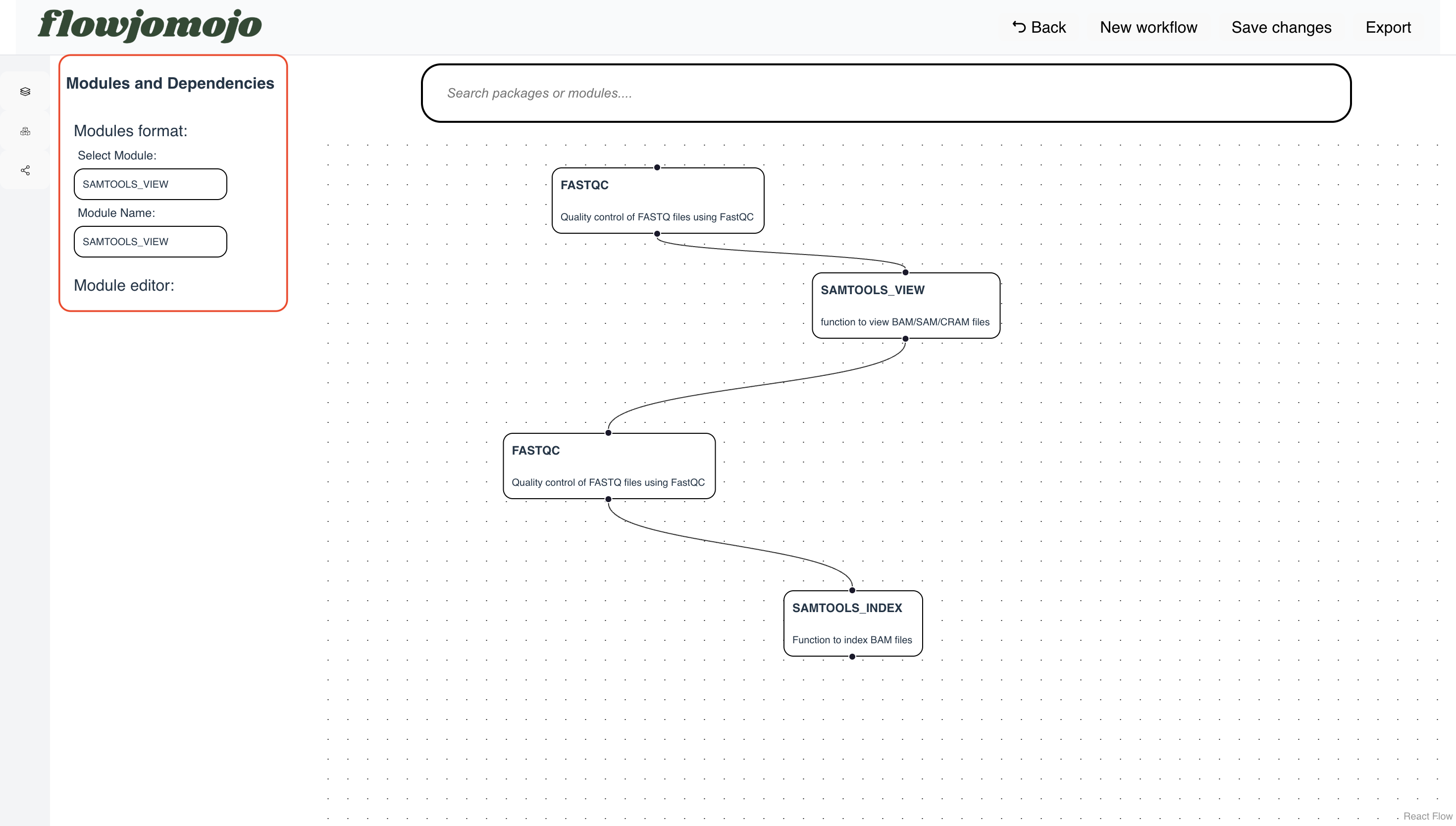
Flowjomojo - React-Flow webapp to construct modules drag and drop for pipeline construction
Tools:
ReactFlow
TypeScript
Abstract:
Flow-based workflow generation from the comfort of your browser to help you keep your flow, Jo. Flowjomojo is a ReactFlow-based web-app that generates ready-made Nextflow/WDL pipelines from drag-and-drop modules. It helps easing the process of writing bioinformatics pipelines, provide configuration settings and visualization of workflows.
Flow-based workflow generation from the comfort of your browser to help you keep your flow, Jo. Flowjomojo is a ReactFlow-based web-app that generates ready-made Nextflow/WDL pipelines from drag-and-drop modules. It helps easing the process of writing bioinformatics pipelines, provide configuration settings and visualization of workflows.
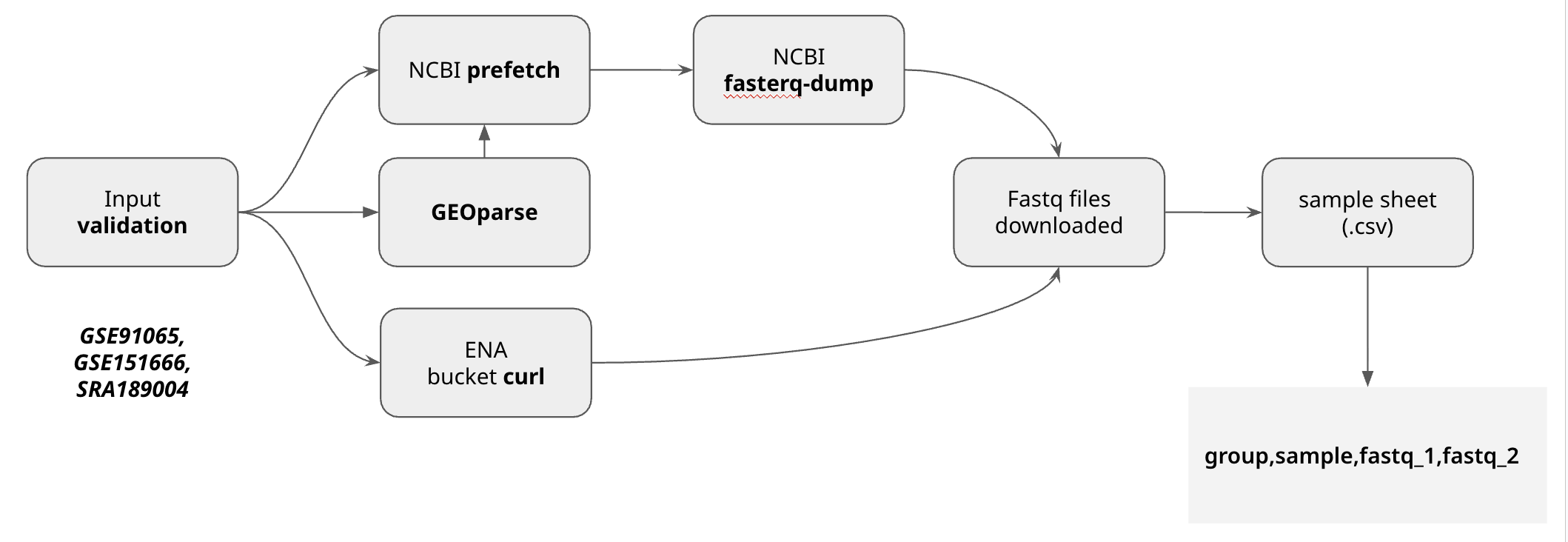
Biofetch - A Nextflow pipeline to parallely retrieve sequencing data from public sites
Tools:
nextflow
sra-toolkits
Abstract:
Biofetch automatically download the fastq files from your samplesheet csv, as long as they have valid accession ID Available sources for retrieval:
- ENA
- SRA
- GEO
Biofetch outputs a samplesheet ready for running your next pipeline.
Biofetch automatically download the fastq files from your samplesheet csv, as long as they have valid accession ID Available sources for retrieval:
- ENA
- SRA
- GEO
Biofetch outputs a samplesheet ready for running your next pipeline.